Data DOI: 10.15785/SBGRID/187 | ID: 187
Kirchhausen Laboratory, Children's Hospital Boston
Release Date: 16 Oct 2015
1. If this dataset is locally available, it should be accessable at /programs/datagrid/187
2. To download this dataset, please run the following command from your Terminal on a Linux or OS X workstation:
'rsync -av rsync://data.sbgrid.org/10.15785/SBGRID/187 .' (Harvard Medical School, USA)
Depending on your location, faster access may be available from a Tier 1 site closer to your location
'rsync -av rsync://sbgrid.icm.uu.se/10.15785/SBGRID/187 .' (Uppsala University, Sweden)
'rsync -av rsync://sbgrid.pasteur.edu.uy/10.15785/SBGRID/187 .' (Institut Pasteur de Montevideo, Uruguay)
'rsync -av rsync://sbgrid.ncpss.org/10.15785/SBGRID/187 .' (Shanghai Institutes for Biological Sciences, China)
3. After the transfer is completed, please issue the following command to verify data integrity:
'cd 187 ; shasum -c files.sha'
Storage requirements: 26G
Biological Sample:
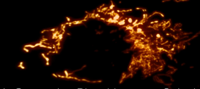
Developing Zebrafish Embryo
Dataset Type:
Lattice Light-Sheet Microscopy
Subject Composition:
DNA, Ligand, Protein, RNA
Collection Facility:
Harvard Medical School - mSlice 1 - Kirchhausen Lab
Data Creation Date:
18 Dec 2014
Related Datasets:
None
Upadhyayula, S; Kirchhausen, T. 2015. "Lattice Light-Sheet Microscopy data for: Developing Zebrafish Embryo.", SBGrid Data Bank, V1, https://doi.org/10.15785/SBGRID/187.
mSlice Conditions: Waveform type : Linear X Galvo Offset, Interval (um), # of Pixels for Excitation (0) : 0 0.1 51 Z Galvo Offset, Interval (um), # of Pixels for Excitation (0) : -0.599257 0 251 Z PZT Offset, Interval (um), # of Pixels for Excitation (0) : 26 0 251 S PZT Offset, Interval (um), # of Pixels for Excitation (0) : 50 0.398406 251 # of stacks (0) : 200 Excitation Filter, Laser, Power (%), Exp(ms) (0) : N/A 488 20 40 Cycle lasers : per Z Z motion : Sample piezo
| Name | Additional Roles | Affiliation While Working on the Project |
|---|---|---|
| Srigokul Upadhyayula | Data Collector, Depositor | Harvard Medical School / BCH |
| Tom Kirchhausen | PI | Children's Hospital Boston |
Deskew: 31.5 degrees geometric transformation to convert raw data into real space. Segmentation Software: ACME: Automated Cell Morphology Extractor for Comprehensive Reconstruction of Cell Membranes; DOI: 10.1371/journal.pcbi.1002780
License: CC0
Terms: Our Community Norms as well as good scientific practices expect that proper credit is given via citation. Please use the data citation, as generated by the SBGrid Data Bank.